Sister of Sex Lethal – a novel translation repressor protein from Drosophila
Driven by the hypothesis that other proteins might regulate translation by mechanisms analogous to Sxl we set out to biochemically characterize Sxl-related RNA-binding proteins. In particular we are interested in a protein termed Sister of Sex Lethal (Ssx), which originated from a gene duplication event early in Drosophilid evolution. Both proteins still share a high degree of identity and we could demonstrate that they associate with similar RNA elements.
While both proteins are able to repress translation via 5’ UTR binding sites in a uORF-dependent manner, the ability of Ssx to regulate translation via the msl-2 3’ UTR is severely compromised. This now allows us to dissect the requirements of this regulatory pathway on a molecular level.
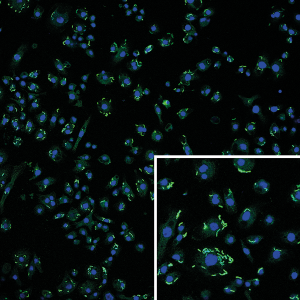
Knockout of Ssx in flies results in hypersensitivity to infection with grampositive bacteria, suggesting a function of Ssx in innate immunity. In order to study the role of Ssx in bacterial infection, we have teamed up with the lab of Ana Eulalio (University of Würzburg) to set up a tissue culture system to study infection of cultured Drosophila cells by Listeria monocytogenes (as shown below, nuclei of the Drosophila cells in blue, Listeria in green).