RNA-binding proteins in Drosophila development and sexual differentiation
Post-transcriptional regulation of gene expression is mostly governed by RNA-binding proteins (RBPs) that control all aspects of RNA biology. In Drosophila melanogaster, female development is governed by a single RNA-binding protein, Sex-lethal (Sxl), that controls the expression of key factors involved in sex-specific differences in morphology, behavior, and dosage compensation.
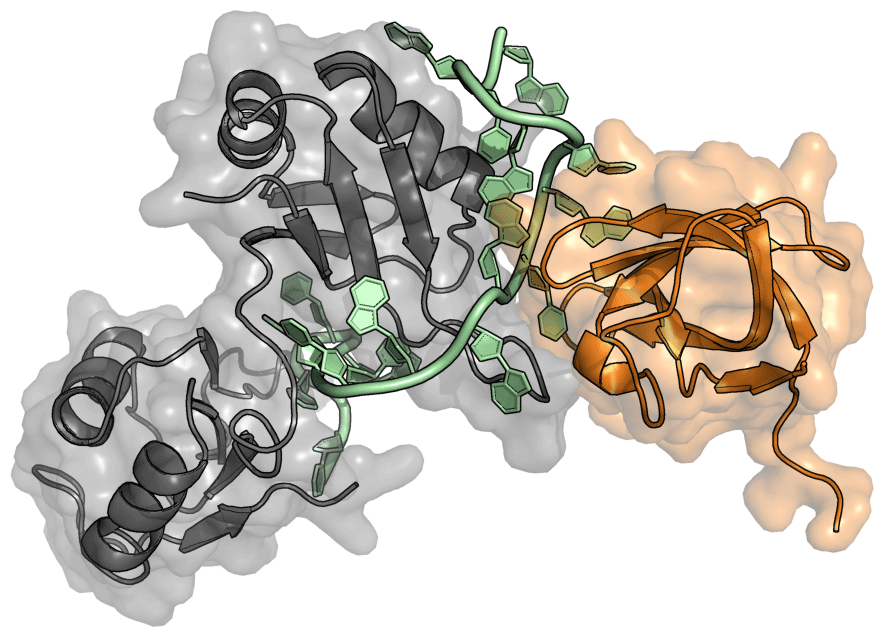
Using Sex-lethal and related proteins as a model system, we aim to understand in molecular detail how RBPs exert their function. Despite its rather simple architecture, Sxl can control gene expression on multiple levels from regulation of alternative splicing and polyA-site choice to translation. To exert function, it often relies on the recruitment of additional factors. However, in most cases the details of how Sxl orchestrates assembly of ribonucleoproteins and how regulation is exerted remain only poorly understood.
Currently we are focusing on:
- the role of Sxl in controlling the switch between mitosis and meiosis in the female germline,
- elucidating the function of the Sex-lethal-related protein Sister-of-Sex-lethal (Ssx), and
- improving the methodology to analyze RBP function in select tissues of living animals
Timeline and major achievements
Select publications
Medenbach, J., Seiler, M., and Hentze, MW.: Translational control via protein-regulated upstream open reading frames. Cell, 2011, 145(6):902-13, https://doi.org/10.1016/j.cell.2011.05.005
Moschall R, Rass M, Rossbach O, Lehmann G, Kullmann L, Eichner N, Strauss D, Meister G, Schneuwly S, Krahn M, and Medenbach J: Sister of Sex-lethal antagonizes Sxl-dependent alternative splicing to maintain a male-specific gene expression pattern in Drosophila Nucleic Acids Res. Nucleic Acids Res., 2019, 18;47(5):2276-2288, https://doi.org/10.1093/nar/gky1284
Moschall R, Strauss D, Garcia-Beyaert M, Gebauer F, Medenbach J: Drosophila Sister-of-Sex-lethal is a repressor of translation. RNA, 2018, 24:149-58, https://rnajournal.cshlp.org/content/24/2/149.full
Müller-McNicoll M, Rossbach O, Hui J, and Medenbach J: Auto-regulatory feedback by RNA-binding proteins – Journal of Molecular Cell Biology 2019, https://doi.org/10.1093/jmcb/mjz043
Moschall R, Gaik M, Medenbach J.: Promiscuity in post-transcriptional control of gene expression: Drosophila Sex-lethal and its regulatory partnerships. FEBS Lett. 2017, 591(11):1471-1488, https://doi.org/10.1002/1873-3468.12652
Salerno-Kochan A, Horn A, Ghosh P, Nithin C, Kościelniak A, Strauss D, Rossbach O, Bujnicki JM, Gaik M, Medenbach J, Glatt S: Molecular insights into RNA recognition and gene regulation by the TRIM-NHL protein Mei-P26. bioRxiv, 2021, https://doi.org/10.1101/2021.09.20.461029
Urdaneta EC, Vieira C , Hick T, Wessels HH, Moschall R, Medenbach J, Ohler U, Granneman S, Selbach M, and Beckmann BM: Purification of Cross-linked RNA-Protein Complexes by Phenol-Toluol Extraction. Nat Commun, 2019, 1;10(1):990, https://doi.org/10.1038/s41467-019-08942-3
Collaborating labs
Sebastian Glatt, Max Planck Research Group at the Malopolska Centre of Biotechnology/Jagiellonian University, Krakow, Poland
Oliver Rossbach, Institute of Biochemistry, Justus-Liebig-University Giessen, Germany
Stephan Schneuwly, Developmental Biology, University of Regensburg, Germany
Benedikt Beckmann, Integrative Research Institute (IRI) for the Life Sciences, Humboldt Universität, Berlin, Germany
Albrecht Bindereif, Institute of Biochemistry, Justus-Liebig-University Giessen, Germany
Fatima Gebauer, Centre for Genomic Regulation (CRG) , Barcelona, Spain
Funding
We are thankful for the financial support by the following agencies:















