Gene regulation during cellular stress and its role in disease and resistance to therapy
Cells are exposed to various environmental stresses and have evolved dedicated genetic programs to survive and proliferate despite the challenges. Cellular stress response pathways have emerged as drivers of human diseases including cancer, diabetes and neurodegeneration. We are using multi-omics analyses to gain insight into post-transcriptional gene regulation during stress.
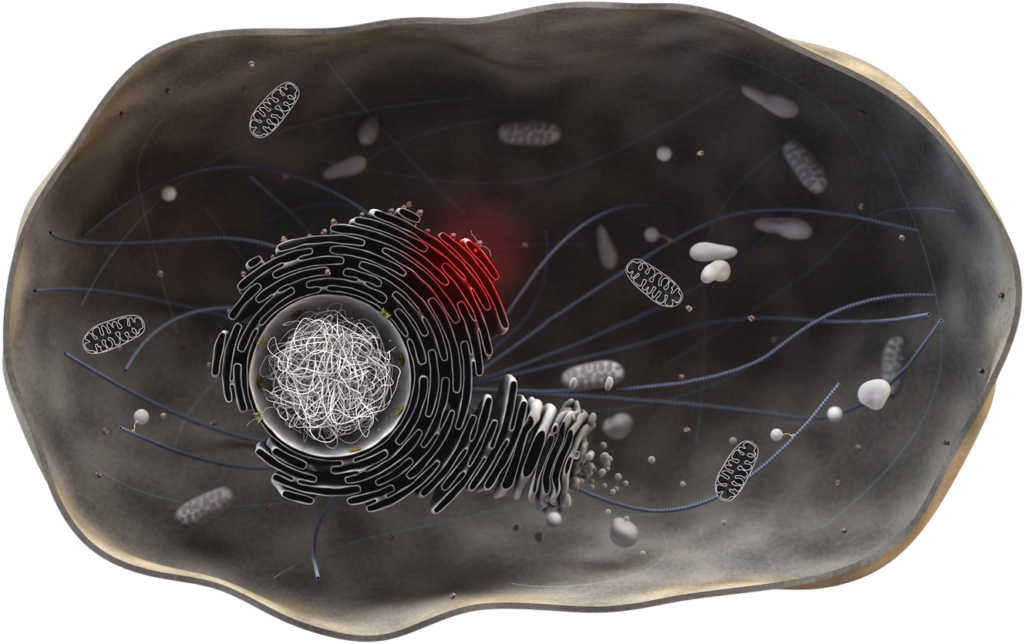
The Unfolded Protein Response (UPR)
The endoplasmic reticulum (ER) is the main cellular compartment in protein folding and maturation with about one third of the proteome being synthesized at the ER. Perturbation of ER proteostasis and accumulation of unfolded proteins trigger an adaptive signal transduction pathway, the Unfolded Protein Response (UPR), that adjusts gene expression to restore cellular homeostasis.
To gain a better understanding of the UPR we have generated comprehensive datasets of the UPR in cellular models using a multi-omics approach. This endeavor was pursued in a collaborative effort with the labs of R. Ahrends (ISAS Dortmund), B. Tews (DKFZ Heidelberg), G. Tödt (EMBL Heidelberg), and C. Knobbe-Thomsen (Heinrich-Heine University Düsseldorf) in the framework of the SUPR-G consortium (Systems biology of the Unfolded Protein Response in Glioma) which was funded by the German Federal Ministry of Education and Research (BMBF) in the framework of the e:Med initiative.
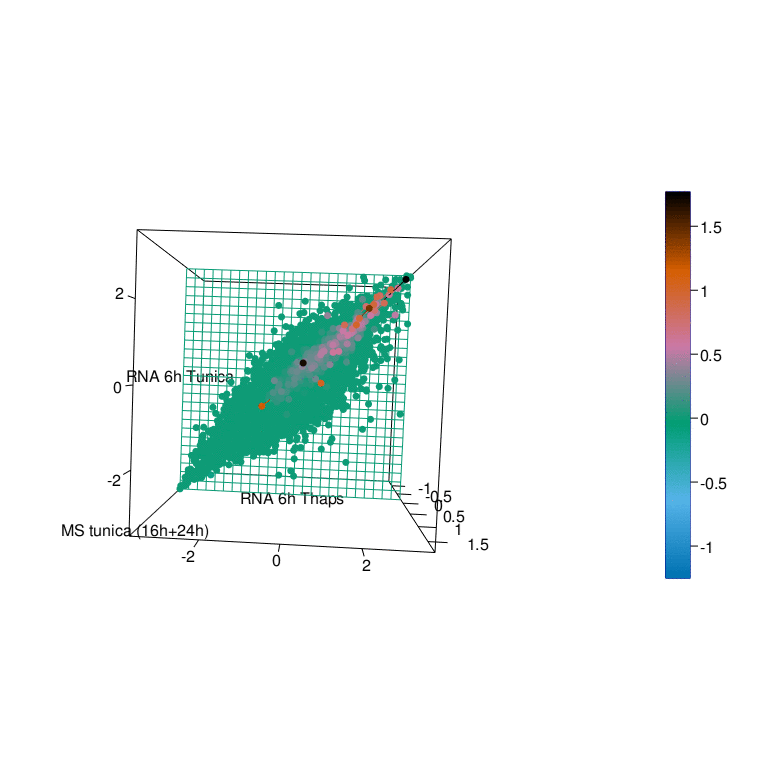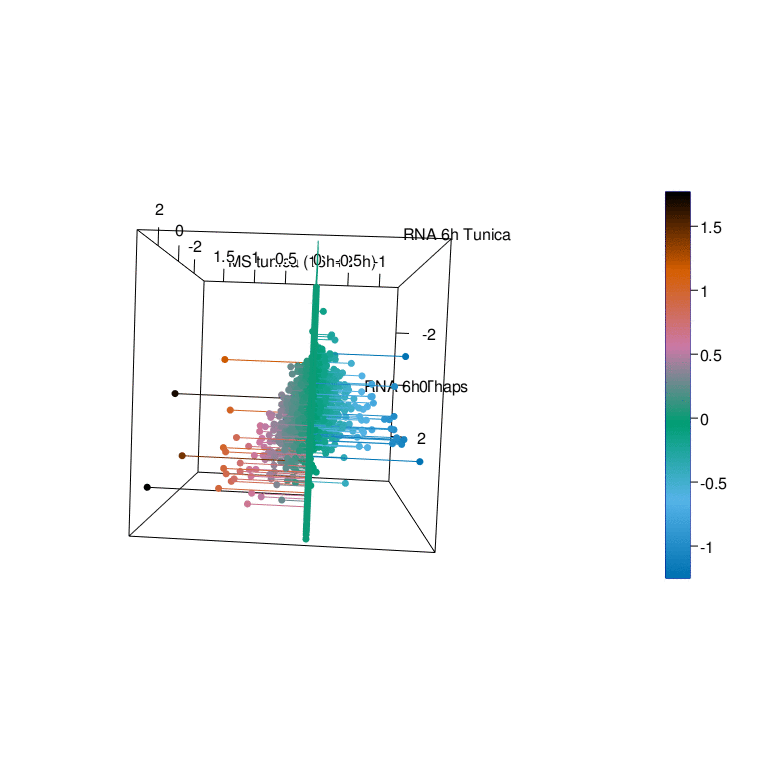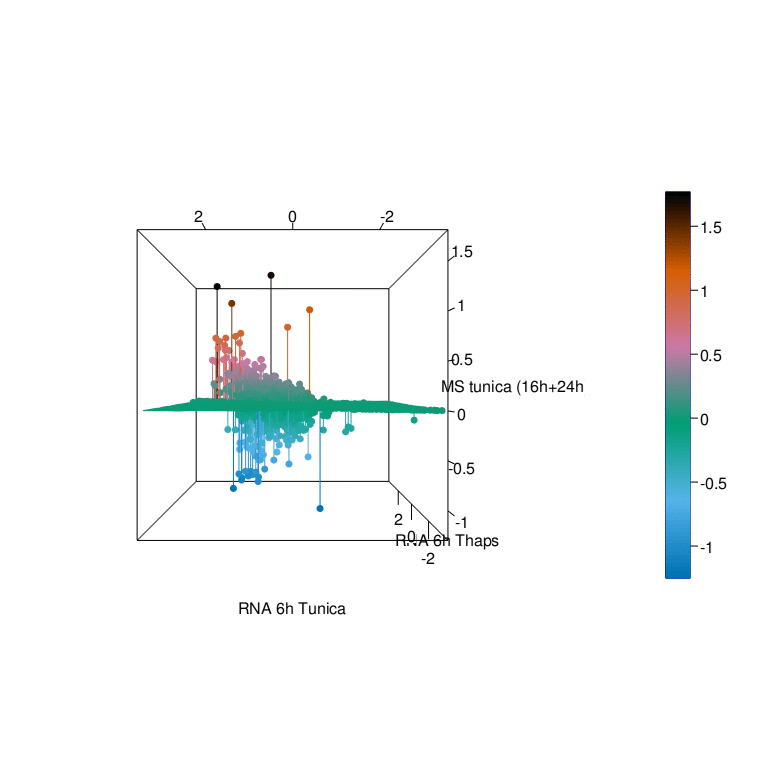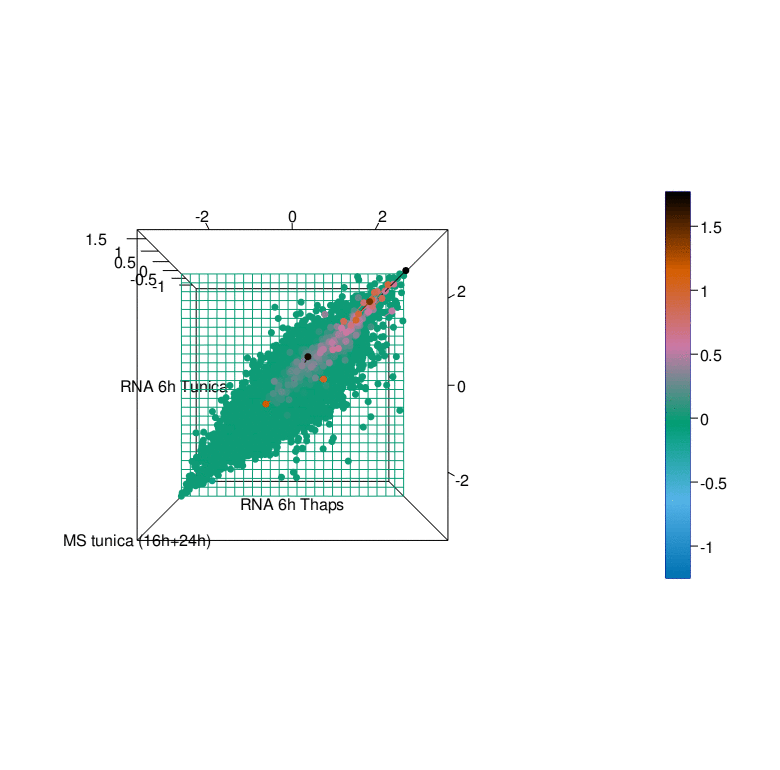
Timeline and major achievements
Select publications
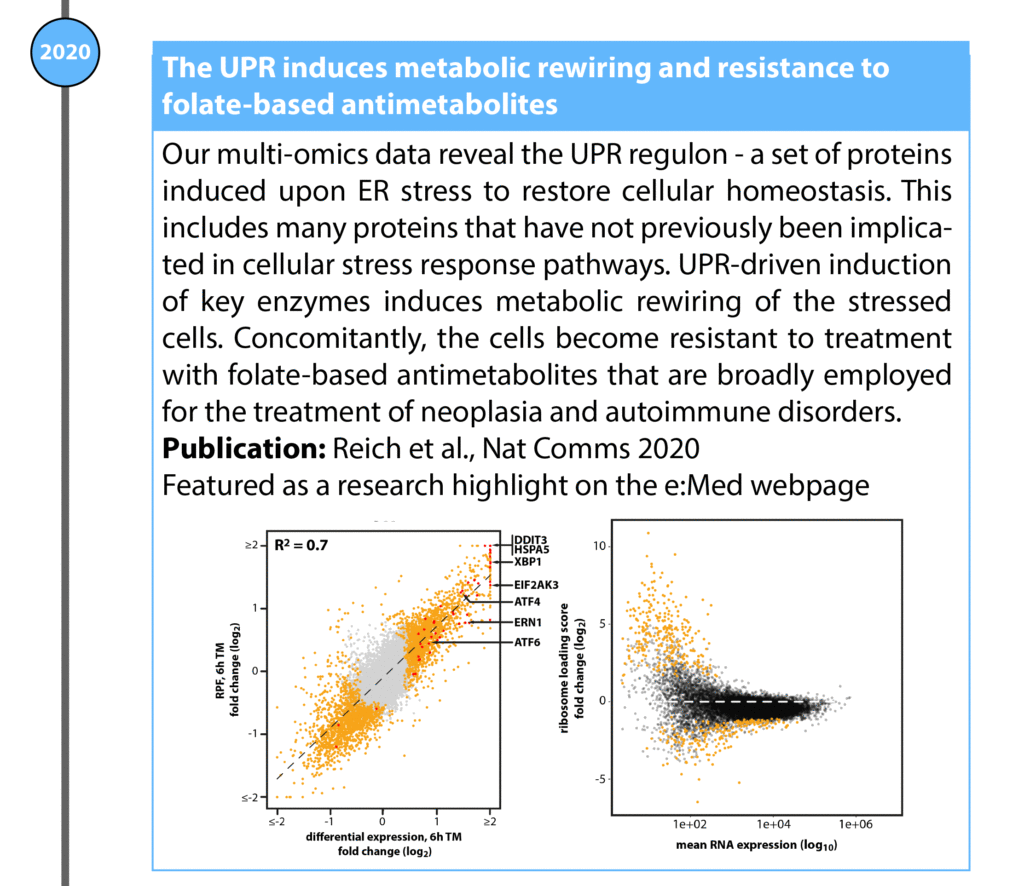
Reich S, Nguyen CDL, Has C, Steltgens S, Soni H, Coman C, Freyberg M, Bichler A, Seifert N, Conrad D, Knobbe-Thomsen CB, Tews B, Toedt G, Ahrends R, and Medenbach J: A multi-omics analysis reveals the unfolded protein response regulon and stress-induced resistance to folate-based antimetabolites – in Nature Communications, DOI:10.1038/s41467-020-16747-y
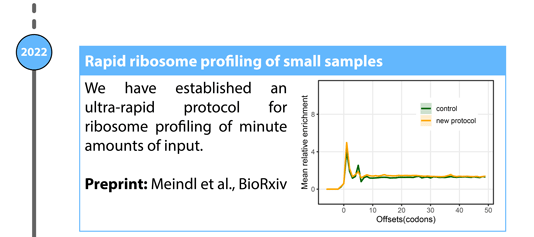
Meindl A, Romberger M, Lehmann G, Eichner N, Kleemann L, Wu J, Danner J, Boesl M, Mesitov M, Meister G, König J, Leidel S, Medenbach J: High resolution and rapid ribosome profiling from low-input samples. bioRxiv, 2022, https://doi.org/10.1101/2022.09.23.509038
Nguyen C, Malchow S, Reich S, Loroch S, Sickmann A, Tews B, Medenbach J, and Ahrends R: PRM-deep: A sensitive and simple targeted proteomics approach to quantify transcription factor and membrane proteins of the unfolded protein response pathway in glioblastoma cells. Sci. Rep., 2019, 9, Article number: 8836, https://doi.org/10.1038/s41598-019-45237-5
Kopczynski D, Hentschel A, Coman C, Schebb NH, Hornemann T, Mashek DG, Hartung NM, Shevchuk O, Schött H-F, Lorenz K, Torta F, Burla B, Zahedi RP, Sickmann A, Kreutz MR, Ejsing CS, Medenbach J, and Ahrends J: Simple targeted assays for metabolic pathways and signaling: a powerful tool for targeted proteomics, Analytical Chemistry, accepted for publication. https://doi.org/10.1021/acs.analchem.0c02793
Wirthschaft P, Bode J, Soni H, Dietrich F, Krüwel T, Fischer B, Knobbe-Thomsen CB, Rossetti G, Hentschel A, Mack N, Schönig K, Breckwoldt MO, Schmandke A, Pusch S, Bendszus M, Schwab ME, Medenbach J, von Deimling A, Kool M, Herold-Mende C, Reifenberger G, Ahrends R, and Tews B: RhoA regulates translation of the Nogo-A decoy SPARC in white matter-invading gliomas – Acta Neuropathol (2019), https://doi.org/10.1007/s00401-019-02021-z
Collaborating labs
Robert Ahrends – Department of Analytical Chemistry, University of Vienna (previously at the Leibniz-Institute for Analytical Sciences (ISAS), Dortmund, Germany)
Grischa Tödt – previuosly at the Structural and Computational Biology Unit, European Molecular Biology Laboratory (EMBL), Heidelberg, Germany
Bjoern Tews – previously at the German Cancer Research Center (DKFZ), Heidelberg, Germany, now at AbbVie
Christiane Knobbe-Thomsen – previously at the Heinrich Heine University Düseldorf
Funding
We are grateful for the financial support by the federal Ministry of Education and research (BMBF) in wthe framework of the e:Med initiative.