New manuscript from the lab published in Nucleic Acids Research: Drosophila Sister-of-Sex-lethal reinforces a male-specific gene expression pattern by controlling Sex-lethal alternative splicing.
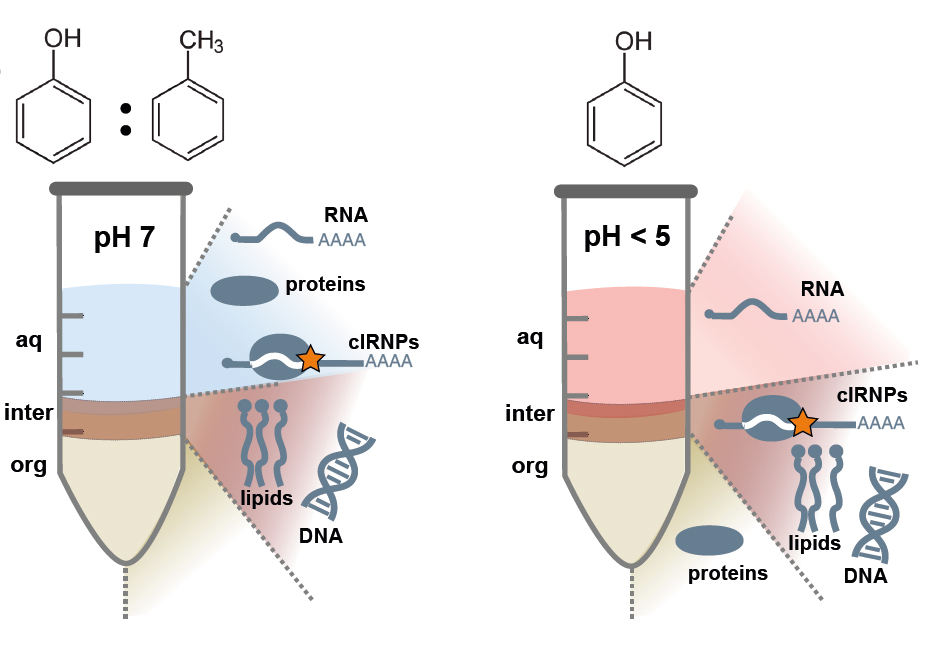
In a collboration with the labs of Stefan Schneuwly, Gunter Meister (both at the University of Regensburg), Michael Krahn (Westfälische Wilhelms-Universität Münster), and Oliver Rossbach (Justus-Liebig-University Giessen), we could demonstrate that the protein Sister-of-sex-lethal (Ssx) is required in male flies to suppress production of Sex-lethal (Sxl).
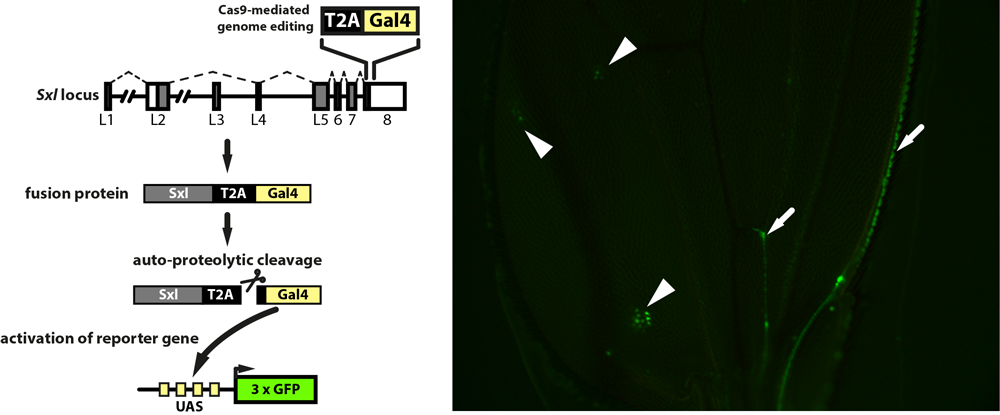
Genomic tagging of the Sex-lethal (Sxl) locus in flies to reveal Sxl protein mis-expression (arrowheads) in male flies mutant for Sister-of-Sex-lethal. Arrows mark expression of a Sxl isoform in neural cell bodies and projections.
Most higher eukaryotes reproduce sexually, increasing the variability in the offspring. This allows e.g. rapid adaption to a new (or changing) environment or the cleansing of harmful mutations from a population. Sexual reproduction in higher eukaryotes usually involves individuals of different sex: males and females. Not surprisingly, the genetic programs that determine sex and control sexual differentiation need to be particularly robust in order to ensure survival of the population.
In Drosophila, a single protein, the master regulator Sex-lethal (Sxl), governs female development by controlling the expression of key factors involved in female morphology and behaviour. Once expressed, it engages in an auto-regulatory, positive feedback loop to ensure its sustained expression. This stably ‘flips the switch’ and commits to female development.
In contrast, in males Sxl expression needs to be shut-off which is achieved by alternative splicing that generates RNA isoforms encoding truncated, non-functional Sxl protein. Fluctuations inherent to gene expression can, however, produce small amounts of functional Sxl protein. When left unchallenged, this protein can trigger a self-enforcing cascade resulting in Sxl protein expression snowballing out of control. Until now, however, it remained unclear how males completely shut off the Sxl expression cascade and protect themselves against runaway protein production to ensure robust sex-specific development.
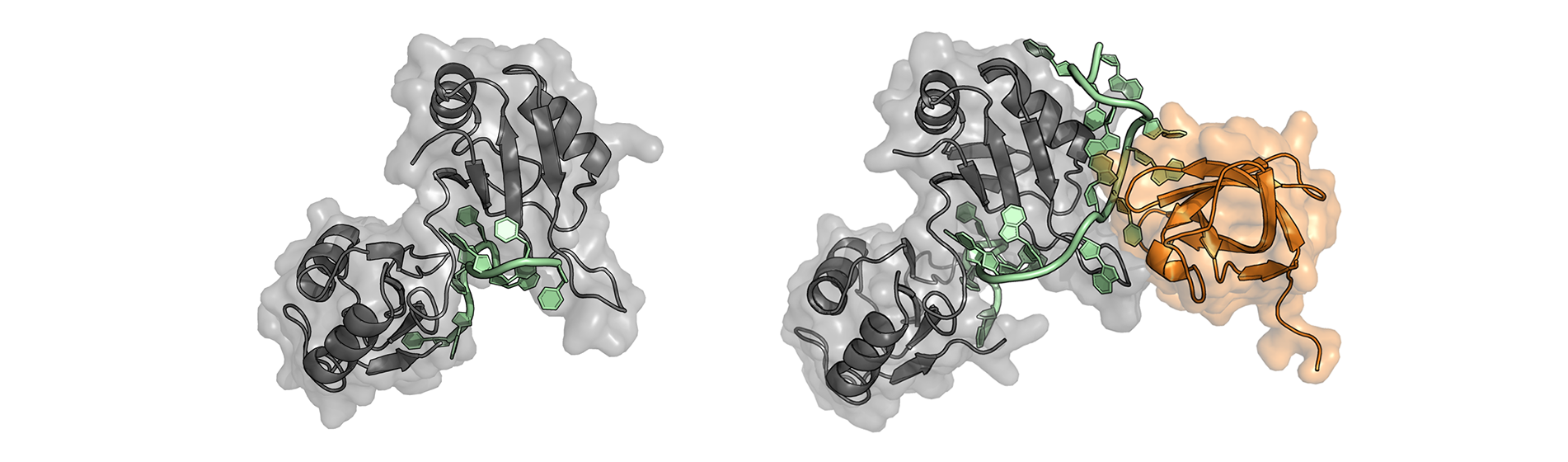
We have discovered a safeguard mechanism that prevents Sxl production in adult male flies. We identified the protein Sister of Sex-lethal (Ssx) as the first antagonist of Sxl-mediated auto-regulatory splicing that defines a precise threshold level for activation of the auto-regulatory, positive feedback loop that controls Sxl expression. We could show that Ssx exerts function by competing with Sxl for the same RNA regulatory elements thus preventing Sxl from triggering the self-enforcing expression cascade in adult male animals.
 Principal Investigators third funding period (from left to right): J. Medenbach, J. Griesenbeck, T. Heise, P. Milkereit, W. Seufert, A. Bruckmann, S. Ferreira-Cerca, M. Kretz, J. Perez-Fernandez, T. Dresselhaus, G. Längst, R. Sprangers, H. Tschochner, D. Grohmann, and C. Engel;
Principal Investigators third funding period (from left to right): J. Medenbach, J. Griesenbeck, T. Heise, P. Milkereit, W. Seufert, A. Bruckmann, S. Ferreira-Cerca, M. Kretz, J. Perez-Fernandez, T. Dresselhaus, G. Längst, R. Sprangers, H. Tschochner, D. Grohmann, and C. Engel;